Despite being over 15 years old, GlycoWorkbench is a mainstay for the field of glycomics. The ability to predict and annotate fragments for glycan structures drawn in a GUI is a feature yet to be matched by other software. Due to its age, Glycoworkbench isn’t compliant with current glycobiology standards, namely the Symbol Nomenclature for Glycans (SNFG). Fortunately, the developers built a method to update your GlycoWorkbench to draw structures and annotate fragments in an SNFG-compliant manner. Here’s a guide on how to do it for yourself.
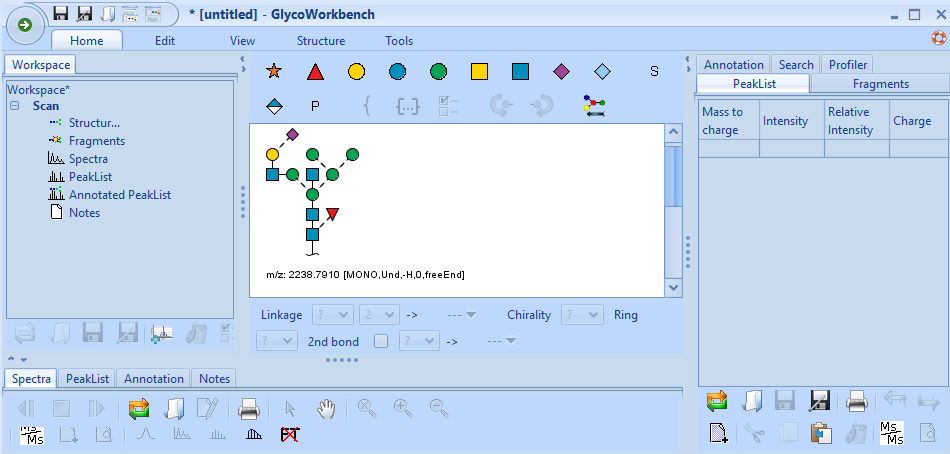
Here is how to do it:
- Download the necessary files to update GlycoWorkbench at the following link:
https://drive.google.com/drive/folders/1qR1fO-opYgU9jLanzN772Sdp1Jikrtl9?usp=drive_link - Click on the green arrow, and then click on “Settings”. Your settings should now appear
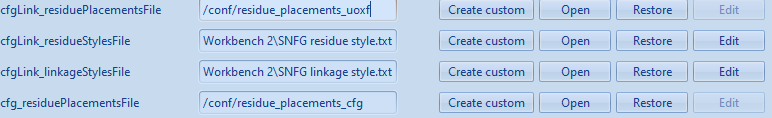
- Scroll down until you find a settings beginning with “cfgLink_”, we will replace those with the current SNFG format. To do so, click the Open button next to “cfgLink_residueStylesFile” and select the recently downloaded file, “SNFG residue style.txt”. Then do the same for “cfgLink_linkageStylesFile”, selecting the recently downloaded file, “SNFG linkage style.txt”.
It should now look like this:
- Click on the X of the Settings window to close it, then click the X on the GlycoWorkbench application to close it as well. Reopen GlycoWorkBench and click on the View tab. Select the middle icon and now your glycans will all be up-to-date with the current publication standards for drawing glycans.
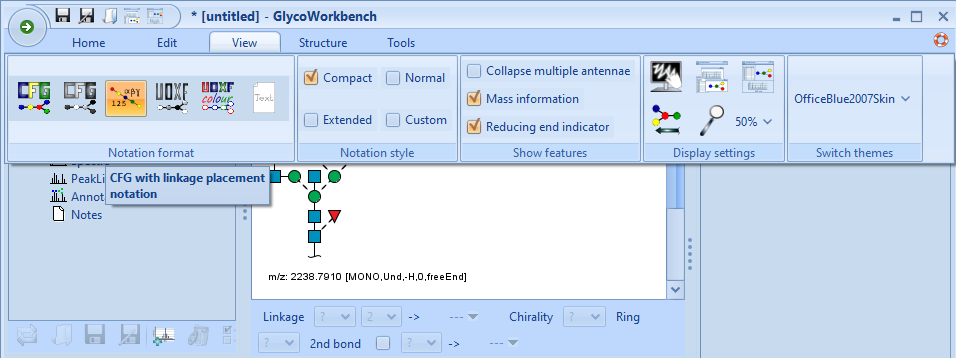
You should now have your glycans drawn in the correct format, eliminating the need for manually drawing glycans in illustrator or other applications. If you have any issues, feel free to contact me at chris (at) proteaglyco.com or Twitter (@proteaglyco). Also, if there is a tutorial you’d like to see, feel free to contact me for that too!
